Cardiovascular diseases
Cardiac image analysis
Cardiac imaging has experienced an impressive increase in quality and uptake in clinical routine. Current CT scanners are able to acquire full cardiac volumes in one rotation, leading to high-quality 4D (3D+time) CT datasets. This in turn leads to the need for dedicated software tools to automatise the analysis of these large quantities of image data, which is one of our focus areas in cardiovascular research.
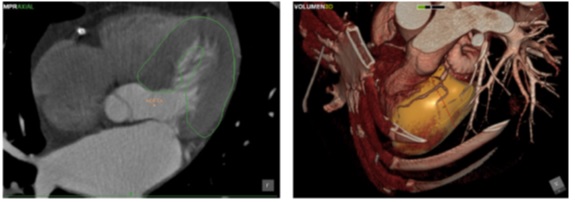
Framework for automatic heart segmentation based on active shape models (Perez et al., Computer Methods and Programs in Biomedicine 2013).
MRI technology is still the gold standard for soft tissue imaging, not least for the depiction of subtle tissue modifications due to heart disease. Cardiac scar tissue, related to infarcts, can be seen in contrast-enhanced MRI images. However, the contrast of the scar and neighbouring tissue is very difficult to perceive, difficulting the establishment of solid quantitative criteria for tissue characterisation. We have developed algorithms for scar detection and segmentation, leading to accurate measures of tissue damage.
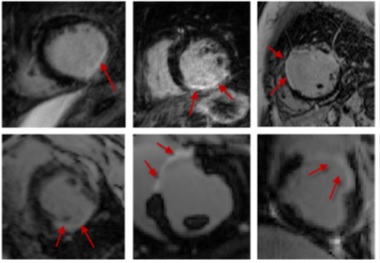
Cardiac scar segmentation based on support vector machines and level sets (Karim et al., Medical Image Analysis 2016)
Cardiac movement analysis
The advent of multiple imaging modalities and longitudinal follow-up acquisitions leads to the need for methods for image registration. Registration is the process of aligning two images so their information can be fused or compared. The researchers in our group have a long track record on the development of theoretical and practical approaches for registration. in particular, cardiac temporal sequences can be registered to model cardiac movement, using methods for deformable registration.
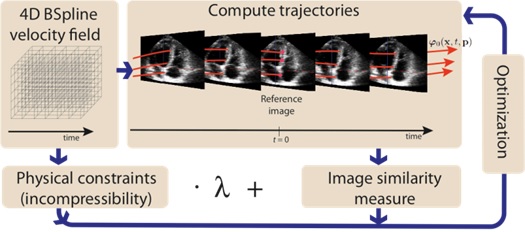
Registration via Temporal Diffeomorphic Free Form Deformations - TDFFD (Piella et al., Medical Image Analysis 2013)
The analysis of motion patterns can be further explored by machine learning techniques. Statistical classifiers and manifold learning methods are developed in our group from a theoretical standpoint, and then applied to the analysis of cardiac motion. Recent work also considers the use of clinical data together with the images, requiring the need for techniques dealing with inhomogeneous data sources and missing data.
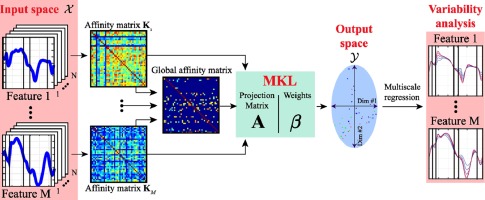
Characterization of myocardial motion patterns by unsupervised multiple kernel learning (Sanchez et al., Medical Image Analysis 2017)
Atherosclerosis
Focusing on vascular diseases, a multiphysics simulation model of atheroma plaque formation was developed. This model, based on Agent-Based simulations, represents the complex processes leading to atherosclerosis, a vascular disease of high prevalence and fatal consequences.
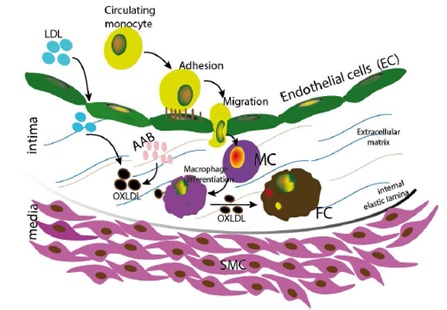
Atherosclerosis formation model, used for agent-based simulations of plaque formation (Olivares et al., Bioinformatics 2016)
Aneurysms
Abdominal aortic aneurysms can be treated via implantable stents, with the help of X-ray fluoroscopic intraoperative guidance. This procedure, however, has severe limitations in regards to the visual information that the interventional radiologist has access to. Typically, the catheter that is used to deploy the stent is visible, but the blood vessels can only be seen at certain stages by the injection of contrast agent. To assist this procedure, we develop surgical planning and guidance systems, via 2D-3D registration approaches based on graph matching.
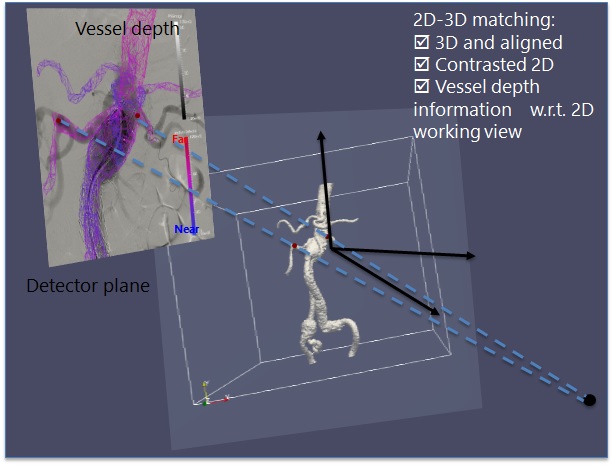
2D-3D registration for intraoperative guidance of stenting procedures for abdominal aortic aneurysms.
