Brain image analysis
Foetal and neonatal brain development
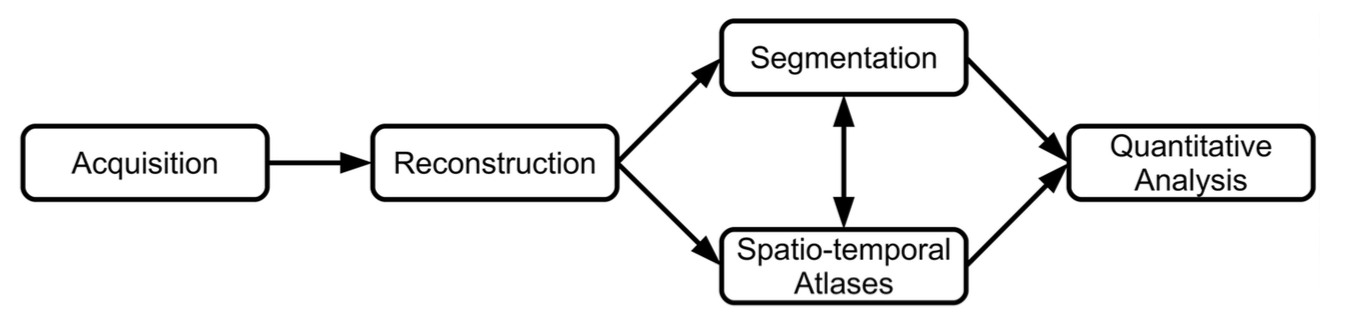
One of our application areas of interest concerns infant brain development. We presented a review of methods for the quantification of fetal brain development (O. M. Benkarim et al. HBM). The pipeline for the analyisis of fetal brains contains several steps.
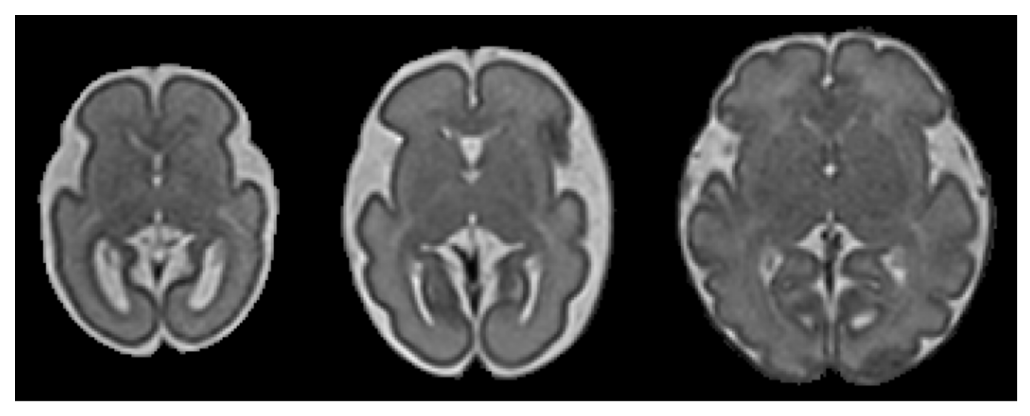
The fetal brain experiences dramatic changes in the later stages of pregnancy. This makes it challenging to obtain stable biomakers characterizing later neurodevelopmental outcomes.
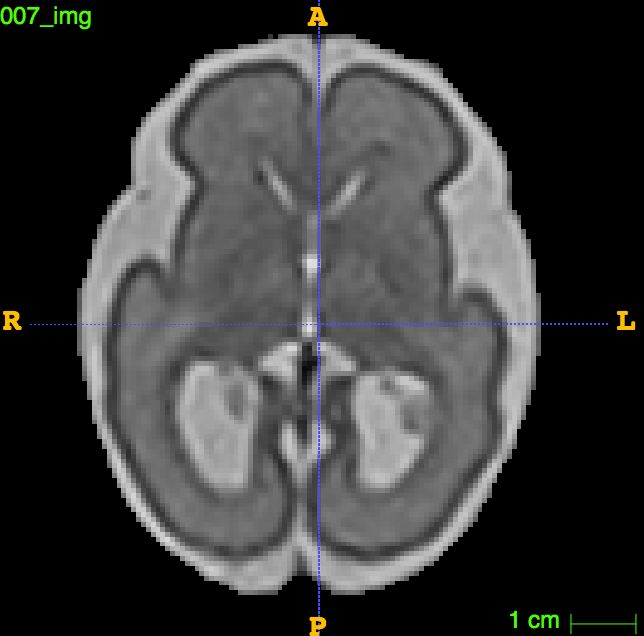
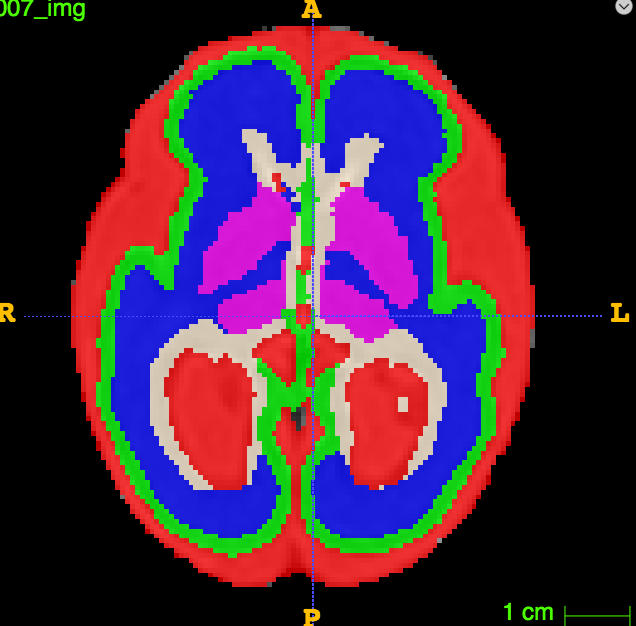
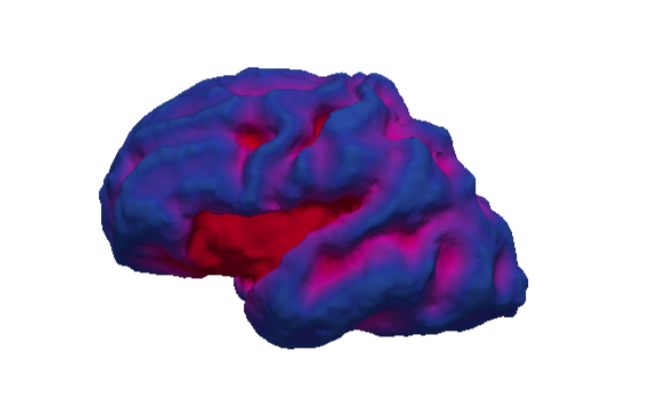
In one of our projects, we want to characterize the differences in cortical growth patterns between fetuses affected with ventriculomegaly and control fetuses. Fetal brains affected by ventriculomegaly are characterized by having enlarged ventricles. Fetal ventriculomegaly is associated to neurodevelopmental problems. The goal of this study is to find predictive biomarkers of later developmental outcomes. To that end, we want to combine morphological features from the fetal brain structures with information from other sources such as gestagional age, birth weight, ...
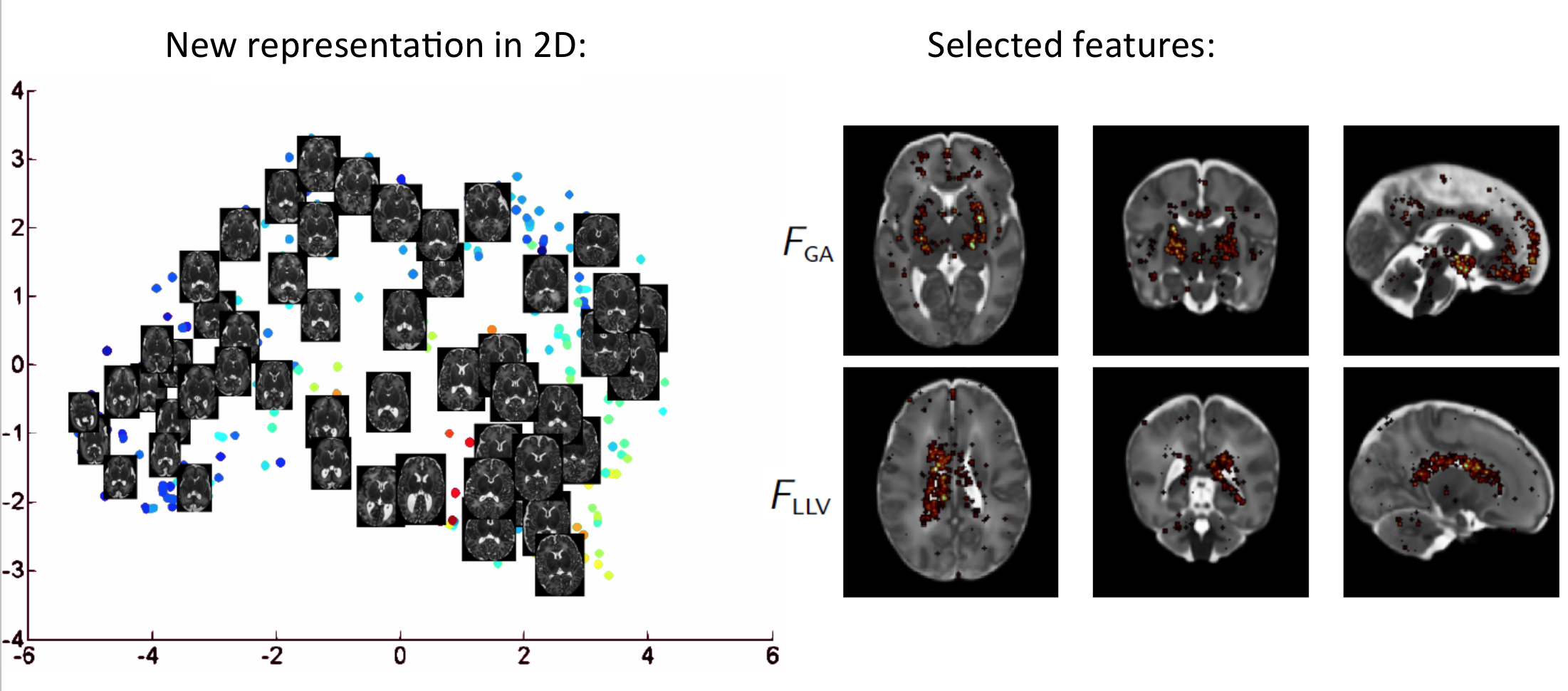
Another line of research concerns the development of computational methods for the analysis of infant brain populations. Our goal is to obtain low-dimensional representations that combine observations from multiple sources such as MRI, diagnostic information, birth weight, gestational age, ... We argue that such low-dimensional representations allow for improved analysis and classification.
Alzheimer's disease
Another area of study concerns neurodegenerative diseases such as Alzheimer's disease (AD). Our projects are divided in two areas 1) development of computational methods capable of handling longitudinal information from multiple modalities and 2) the characterization of the morphological changes related to Alzheimer's disease, more especifically in the region of the hippocampus.
The first project focuses on the development of novel machine learning methods for the early prediction of AD that can specifically handle longitudinal heterogeneous information.
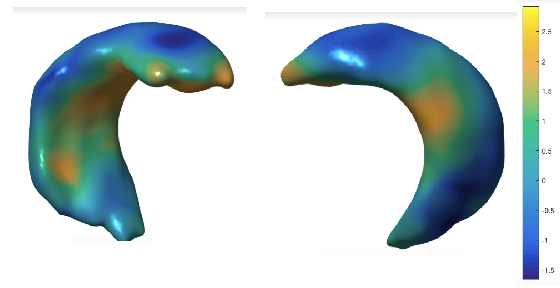
The second project aims at quantifying morphological differences in the hippocampus between populations of interest. In one study, we analyze the effect of the APoE genotype of a healthy on the shape of the hippocampus. The APoE genotype is known to be a risk factor of AD and, unraveling the morphological effects of this genotype might provide insight on the structural changes underlying AD in the incipient stages. We are using the ALFA cohort from Pasqual Maragall Foundation, which provides an excellent opportunity to study the effects of the APoE4 allele due to the high prevalence of APoE4 carriers in their cohort. Below we show the areas in the hippocampus (in yellow) whose shape is affected by the number of APoE4 alleles (0, 1 or 2). These results are obtained with multivariate generalized linear models controlling for the age and gender of the subjects.
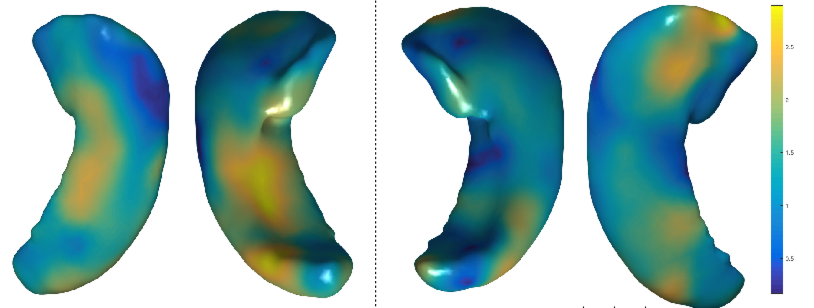
In another study, we aim at quantifying the longitudinal atrophy patterns characteristic of AD. We are using a longitudinal dataset of hippocampal shapes from Alzheimer's transgenic rats from IDIBAPS Institute. Below we show the areas in the hippocampus of the Alzheimer transgenic rats (in yellow) that differ from the normal controls. Results are obtained with linear mixed effects models controling for age and brain volume (fixed effects) and allowing for individual random effects across the multiple measurements of each subject.